Big Data Analytics – Biomedical and Health Informatics for Personalized Cardiovascular Disease Care
By Po-Yen Wu, Janani Venugopalan, Sonal Kothari, Chanchala Kaddi, Chih-Wen Cheng, John H. Phan, and May D. Wang
Advances in “-omics”, devices, imaging, and other advanced biomedical technologies in recent years have resulted in the exponential growth of biomedical data, and have started the era of “Big Data” in health and biomedicine. Cardiovascular Disease is an area where effective use of Big Data analytics hold great promise for risk assessment and personalized medical treatment.
Introduction to Big Data Analytics and Integrative Bioinformatics
Many of these large datasets are available in public repositories such as the Cardiovascular Research Grid and The Cancer Genome Atlas (TCGA). However, the challenge is on how to make sense of these “Big Data” by addressing computational and analytical complexity caused by data volume, velocity, variety, and veracity. To improve personalized health in terms of prediction accuracy, researchers not only have looked into cloud and distributed computing resources for archival and analysis of these huge datasets, but also have started solving analytical complexity by using integrative informatics on multi-modality datasets.
In 2014 Healthcare Information and Management Systems Society (HIMSS’14) annual conference held in Orlando, Florida, cardiovascular disease (CVD) care was presented as an example of pervasive healthcare in the Wearable Sensor and Sensor Informatics Session. CVD care really needs “Big Data” analytics because multiple factors (e.g., genetics, behavior, environment, and clinical courses) are known to play a critical role in disease risk and overall clinical outcome. CVD is the leading cause of death worldwide. In the U.S., though the relative mortality rate has dropped more than 30% in the past decade, CVD still accounts for almost one-third of all deaths in recent years. To effectively reduce the incidence of CVD death and to improve CVD care, it is important to incorporate the concept of “Predictive, Preventive, Personalized, Participatory, Pervasive, Patient-Centric, Pre-emptive, and Precise” (8-P) medicine. The success of 8-P CVD care requires deeper understanding of CVD causes and more accurate projection of CVD events based on identification and integration of CVD molecular biomarkers, imaging markers, and clinical markers.
CVD Biomarker Identification
Microarrays and mass spectrometry are experimental techniques for personalized study of molecular profiles (e.g., gene expression, gene variants, protein activities, and metabolite abundance) in disease diagnosis, prognosis, and treatment. To predict CVD endpoints from microarray data, we have developed a translational bioinformatics pipeline that includes (1) pre-processing, normalization, and quality control, (2) exploratory and unsupervised analysis, (3) feature selection and prediction modeling, and (4) biological interpretation (Figure 1). For microarrays, spatial artifacts due to experimental variation in hybridization chemistry are a major cause of data quality deterioration. Thus, we have developed a web-based bioinformatics tool caCORRECT to detect and remove these artifacts for more accurate gene expression estimates. Once the data have been cleaned, unsupervised exploratory methods such as hierarchical clustering, k-means clustering, and principal component analysis (PCA) may be applied to identify underlying patterns in the data (e.g., patient subgroups and genes with similar expression patterns). Subsequently, supervised feature selection methods may be used to identify differentially expressed genes (DEGs) or biomarkers for clinical endpoint prediction. Comparing numerous feature selection methods, we have shown that knowledge-based feature selection in omniBiomarker can improve prediction performance. We have also shown that using meta-analysis-based biomarker identification from multiple gene expression datasets can further improve prediction performance. In addition, we have conducted a comprehensive microarray data analysis study in the FDA-led MicroArray Quality Control-II (MAQC) consortium. Using one simple algorithm K-nearest neighbor (KNN) as an example, we have shown that a best-practice guideline can lead to better reproducibility in biomarker performance. Once biomarkers are identified, it is critical to validate and interpret the functions of these biomarkers in silico and in the wet-lab. For example, we have designed and developed GOMiner, a Gene Ontology mining tool, to verify that DEGs identified from an atherosclerosis dataset are indeed associated with biological processes of CVD.

Figure 1: The translational bioinformatics pipeline for identifying biomarkers. (a) Quality control: detecting, marking, and compensating for spatial artifacts in microarrays by caCORRECT. (b) Exploratory analysis: hierarchical clustering of samples based on gene expression. (c) Feature selection: discovering discriminant biomarkers using omniBiomarker. (d) Prediction modeling: constructing models that classify samples into given groups. (e) Validation: Q-IHC provides semi-automatic segmentation, morphological feature analysis and quantification, color-based region detection, automatic cell counting, and quantitative molecular profiling for analyzing quantum dot-enhanced immunohistochemical images.
Next-generation sequencing (NGS) technology generates high-resolution, low-cost, and high-throughput base-pair information without assuming a prior knowledge. Among NGS, the whole genome sequencing can identify genome-wide variations, including single nucleotide polymorphisms (SNPs), insertions and deletions, and genomic structural variations ; while the transcriptome sequencing (a.k.a. RNA-sequencing) can profile gene expression, detect gene fusions, identify alternative splicing, and determine SNP expression. Raw NGS base-pair data require multiple analytical steps such as mapping, quantification, normalization, and DEG detection before the translational pipelines in Figure 1. Thus, dozens of bioinformatics solutions have been developed for each of these analytical steps. For example, has investigated the impact of many possible genome annotations on downstream analysis and has identified the importance of the genome annotation for RNA-sequencing. However, as of now, finding the best practice in constructing robust NGS DEG analysis pipelines remains an open research challenge, and still requires extensive research.
CVD Imaging Marker Identification and Integration with Biomarkers
Besides biomarkers, organ and microscopic imaging are clinical diagnostic tools applicable to CVD. For example, organ imaging techniques such as magnetic resonance imaging (MRI), computed tomography, and echocardiogram measure myocardial strain and deformation. Researchers have developed image analysis methods to find cardiac MRI imaging markers for accurate cardiac motion tracking. On the cellular level, microscopic imaging methods can assess inflammation caused by stents or transplants. Automatic analysis of these microscopic images for clinical decision support is an emerging and fast growing area in biomedical imaging informatics. In recent years, we have developed comprehensive tissue biopsy imaging informatics systems that can automatically segment tissue structures, extract imaging descriptors or imaging markers, and predict disease subtypes and disease grades for diagnosis or prognosis (Figure 2 (a)). This system has worked well with cardiac pathological images
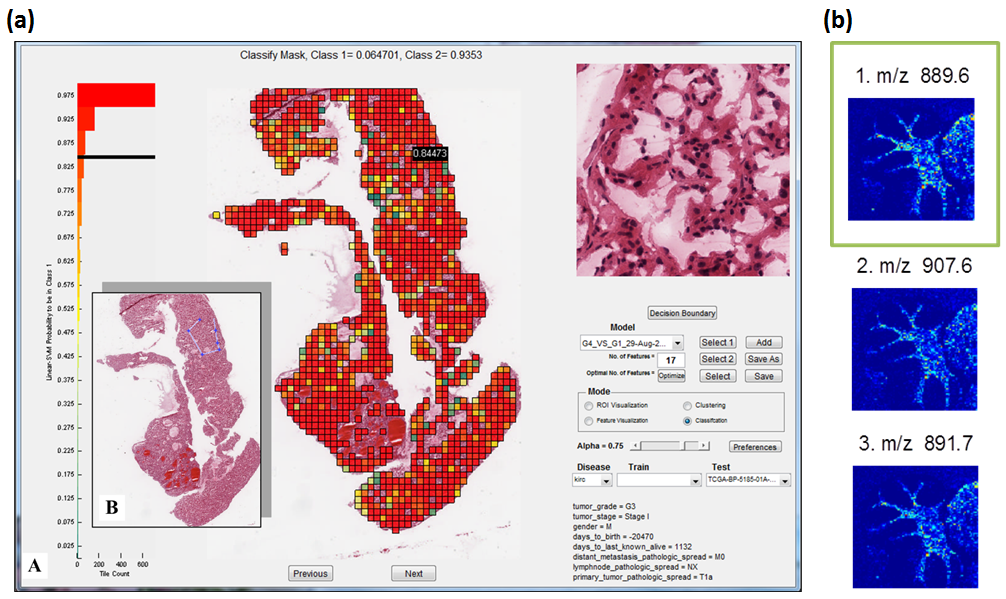
Figure 2: (a) Histopathological whole-slide imaging (WSI) marker identification and clinical decision support: TissueViz. The histogram with label A shows the probability of a region in a WSI being low disease grade in contrast with being high grade. The panel with label B is the original sample with an incomplete polygon showing the manual annotation process. (b) Application of the multivariate hypergeometric similarity measure to identify m/z images similar to a query (outlined) in MSI data.
Moving forward beyond single imaging modality alone, advanced biotechnology such as mass spectrometry imaging (MSI) extends beyond conventional mass spectrometry. MSI measures both the molecular profile and the morphological profile of a tissue biopsy simultaneously. Thus, it can study conditions involving spatially localized biochemical changes or damage such as myocardial infarction. Because each pixel in the biopsy image is represented by thousands of molecules in a mass spectrum, identifying molecular signatures (i.e., biomarkers) requires a translational pipeline similar to that presented in Figure 1 and identifying the spatial pattern (i.e., imaging markers) requires imaging informatics in Figure 2 (a). Up to now, analyzing huge amounts of MSI datasets to identify biologically meaningful molecules or spatial patterns still remains an open challenge. As a first step towards addressing this challenge, we have developed the following two analytics: (1) to perform a knowledge-based query using a similarity measure; that is to identify molecules (in MSI data) or spectra (in both mass spectrometry and MSI data) that are potentially related to a known molecule or spectrum of interest, as shown in Figure 2 (b) ; and (2) to perform exploratory data mining using omniSpect, that is to identify prominent spatial and molecular patterns in MSI data. We hope to use these advanced analytics not only to find the potential biomarkers for CVD, but also to correlate the biomarkers with morphological image markers.
Besides using MSI technology to measure both biomolecules and images simultaneously, another trend is to directly integrate two traditionally independent technologies, e.g., “-omic” and “imaging”, to produce holistic analytical models for disease diagnosis. One major effort in CVD now is to integrate histopathological imaging markers with NGS biomarkers to provide personalized care for cardiac transplant patients.
CVD Clinical Marker Identification
As stated before, 8-P CVD care not only requires more understanding of CVD disease mechanisms enabled by CVD biomarkers and imaging markers, but also requires more advanced data analytics to integrate various types of data relating to CVD. Using the cardiac intensive care unit (CICU) as an example, patients have acute and complex CVD conditions need personalized care. However, because of the rapid growth of all types of measurement technologies in the modern ICU, it has become increasingly challenging to make timely and critical clinical decisions based on the huge amounts of heterogeneous data collected by multi-scale, multi-modality, and multi-velocity sensors. Working with CICU physicians, we have designed and developed a data-driven clinical decision support system, icuARM, using association rule mining (ARM). The system can mine thousands of heterogeneous clinical variables in a clinical setting to provide risk analysis in real-time for decision support (Figure 3). In our case study, clinical markers such as pre-existing comorbidities, medication usage, and demographics are identified as high risk factors for mortality and prolonged ICU length-of-stay.
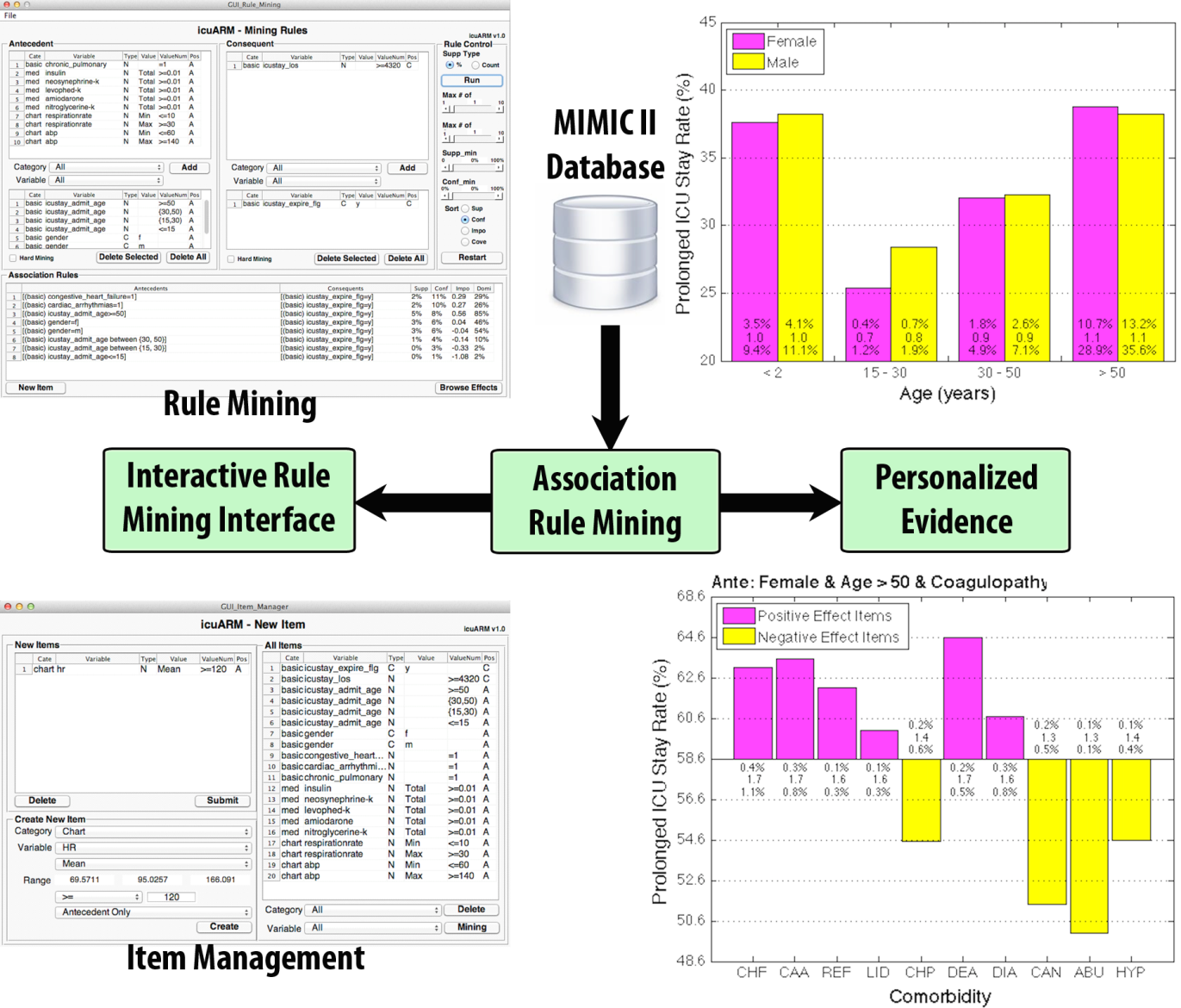
Figure 3: The icuARM system designs ARM rules for mining thousands of clinical factors from heterogeneous datasets in intensive care unit setting.
Summary
In this short overview, we have highlighted the challenges and opportunities of using “Big Data” analytics for personalized CVD care. We have provided evidence that future CVD care will benefit from effective integration of multi-scale, multi-modality, and complex biomedical data through advanced data analytics. Ultimately, “Big Data” analytics can be used to not only assess CVD risk, but also modify therapeutic procedures on a personalized basis to predict treatment outcomes.
Acknowledgements
This work was supported by funding from the National Institutes of Health (NHLBI 5U01HL080711, U54CA119338, and R01CA163256), Georgia Research Alliance Distinguished Cancer Scholar Award to Prof. May D. Wang, and seed grant from Georgia Tech, Emory, and Children’s Healthcare of Atlanta.
For Further Reading
[1} W. Wang and E. Krishnan, “Big Data and Clinicians: A Review on the State of the Science,” JMIR Medical Informatics, vol. 2, p. e1, 2014.[2] R. Winslow, J. Saltz, and I. Foster, “The cardiovascular research grid (cvrg) project,” in AMIA Annu Symp Proc, 2010, pp. 77-81.
[3] V. Marx, “Biology: The big challenges of big data,” Nature, vol. 498, pp. 255-260, 2013.
[4] J. H. Phan, C. F. Quo, C. Cheng, and M. D. Wang, “Multiscale Integration of -Omic, Imaging, and Clinical Data in Biomedical Informatics,” Biomedical Engineering, IEEE Reviews in, vol. 5, pp. 74-87, 2012.
[5] L. A. Cooper, J. Kong, D. A. Gutman, F. Wang, J. Gao, C. Appin, S. Cholleti, T. Pan, A. Sharma, and L. Scarpace, “Integrated morphologic analysis for the identification and characterization of disease subtypes,” Journal of the American Medical Informatics Association, vol. 19, pp. 317-323, 2012.
[6] J. Kong, L. A. Cooper, F. Wang, D. A. Gutman, J. Gao, C. Chisolm, A. Sharma, T. Pan, E. G. Van Meir, and T. M. Kurc, “Integrative, multimodal analysis of glioblastoma using TCGA molecular data, pathology images, and clinical outcomes,” Biomedical Engineering, IEEE Transactions on, vol. 58, pp. 3469-3474, 2011.
[7] G. Thanassoulis and R. S. Vasan, “Genetic Cardiovascular Risk Prediction Will We Get There?,” Circulation, vol. 122, pp. 2323-2334, 2010.
[8] A. S. Go, D. Mozaffarian, V. L. Roger, E. J. Benjamin, J. D. Berry, M. J. Blaha, S. F. Dai, E. S. Ford, C. S. Fox, S. Franco, H. J. Fullerton, C. Gillespie, S. M. Hailpern, J. A. Heit, V. J. Howard, M. D. Huffman, S. E. Judd, B. M. Kissela, S. J. Kittner, D. T. Lackland, J. H. Lichtman, L. D. Lisabeth, R. H. Mackey, D. J. Magid, G. M. Marcus, A. Marelli, D. B. Matchar, D. K. McGuire, E. R. Mohler, C. S. Moy, M. E. Mussolino, R. W. Neumar, G. Nichol, D. K. Pandey, N. P. Paynter, M. J. Reeves, P. D. Sorlie, J. Stein, A. Towfighi, T. N. Turan, S. S. Virani, N. D. Wong, D. Woo, M. B. Turner, A. H. A. S. Comm, and S. S. Subcomm, “Heart Disease and Stroke Statistics-2014 Update A Report From the American Heart Association,” Circulation, vol. 129, pp. E28-E292, Jan 21 2014.
[9] L. Hood and S. H. Friend, “Predictive, personalized, preventive, participatory (P4) cancer medicine,” Nature Reviews Clinical Oncology, vol. 8, pp. 184-187, Mar 2011.
[10] B. A. Stanley, R. L. Gundry, R. J. Cotter, and J. E. Van Eyk, “Heart disease, clinical proteomics and mass spectrometry,” Dis Markersi>, vol. 20, pp. 167-78, 2004.
[11] A. Thomas, S. Lenglet, P. Chaurand, J. Deglon, P. Mangin, F. Mach, S. Steffens, J. L. Wolfender, and C. Staub, “Mass spectrometry for the evaluation of cardiovascular diseases based on proteomics and lipidomics,” Thromb Haemost, vol. 106, pp. 20-33, Jul 2011.
[12] J. H. Phan, C. F. Quo, and M. D. Wang, “Cardiovascular Genomics: A Biomarker Identification Pipeline,” IEEE Transactions on Information Technology in Biomedicine, vol. 16, pp. 809-822, Sep 2012.
[13] R. A. Moffitt, Y. G. Qiqin, T. H. Stokes, R. M. Parry, J. H. Torrance, J. H. Phan, A. N. Young, and M. D. Wang, “caCORRECT2: Improving the accuracy and reliability of microarray data in the presence of artifacts,” Bmc Bioinformatics, vol. 12, Sep 29 2011.
[14] M. B. Eisen, P. T. Spellman, P. O. Brown, and D. Botstein, “Cluster analysis and display of genome-wide expression patterns,” Proceedings of the National Academy of Sciences, vol. 95, pp. 14863-14868, 1998.
[15] F.-X. Wu, “Genetic weighted k-means algorithm for clustering large-scale gene expression data,” BMC bioinformatics, vol. 9, p. S12, 2008.
[16] M. Ringnér, “What is principal component analysis?,” Nature biotechnology, vol. 26, pp. 303-304, 2008.
[17] Y. Saeys, I. Inza, and P. Larrañaga, “A review of feature selection techniques in bioinformatics,” bioinformatics, vol. 23, pp. 2507-2517, 2007.
[18] J. H. Phan, A. N. Young, and M. D. Wang, “omniBiomarker: A Web-Based Application for Knowledge-Driven Biomarker Identification,” IEEE Transactions on Biomedical Engineering, vol. 60, pp. 3364-3367, Dec 2013.
[19] J. H. Phan, A. N. Young, and M. D. Wang, “Robust Microarray Meta-Analysis Identifies Differentially Expressed Genes for Clinical Prediction,” The Scientific World Journal, vol. 2012, 2012.
[20] L. Shi, G. Campbell, W. D. Jones, F. Campagne, Z. Wen, S. J. Walker, Z. Su, T.-M. Chu, F. M. Goodsaid, and L. Pusztai, “The MicroArray Quality Control (MAQC)-II study of common practices for the development and validation of microarray-based predictive models,” Nat Biotechnol, vol. 28, p. 827, 2010.
[21] R. M. Parry, W. Jones, T. H. Stokes, J. H. Phan, R. A. Moffitt, H. Fang, L. Shi, A. Oberthuer, M. Fischer, W. Tong, and M. D. Wang, “k-Nearest neighbor models for microarray gene expression analysis and clinical outcome prediction,” Pharmacogenomics Journal, vol. 10, pp. 292-309, Aug 2010.
[22] B. R. Zeeberg, W. Feng, G. Wang, M. D. Wang, A. T. Fojo, M. Sunshine, S. Narasimhan, D. W. Kane, W. C. Reinhold, and S. Lababidi, “GoMiner: a resource for biological interpretation of genomic and proteomic data,” Genome Biol, vol. 4, p. R28, 2003.
[23] Y. Xing, Q. Chaudry, C. Shen, K. Y. Kong, H. E. Zhau, L. W. Chung, J. A. Petros, R. M. O’Regan, M. V. Yezhelyev, J. W. Simons, M. D. Wang, and S. Nie, “Bioconjugated quantum dots for multiplexed and quantitative immunohistochemistry,” Nat Protoc, vol. 2, pp. 1152-65, 2007.
[24] M. A. DePristo, E. Banks, R. Poplin, K. V. Garimella, J. R. Maguire, C. Hartl, A. A. Philippakis, G. del Angel, M. A. Rivas, M. Hanna, A. McKenna, T. J. Fennell, A. M. Kernytsky, A. Y. Sivachenko, K. Cibulskis, S. B. Gabriel, D. Altshuler, and M. J. Daly, “A framework for variation discovery and genotyping using next-generation DNA sequencing data,” Nature Genetics, vol. 43, pp. 491-+, May 2011.
[25] P. Medvedev, M. Stanciu, and M. Brudno, “Computational methods for discovering structural variation with next-generation sequencing,” Nature Methods, vol. 6, pp. S13-S20, Nov 2009.
[26] Z. Wang, M. Gerstein, and M. Snyder, “RNA-Seq: a revolutionary tool for transcriptomics,” Nature Reviews Genetics, vol. 10, pp. 57-63, Jan 2009.
[27] A. McPherson, F. Hormozdiari, A. Zayed, R. Giuliany, G. Ha, M. G. F. Sun, M. Griffith, A. H. Moussavi, J. Senz, N. Melnyk, M. Pacheco, M. A. Marra, M. Hirst, T. O. Nielsen, S. C. Sahinalp, D. Huntsman, and S. P. Shah, “deFuse: An Algorithm for Gene Fusion Discovery in Tumor RNA-Seq Data,” Plos Computational Biology, vol. 7, May 2011.
[28] P. Y. Wu, J. H. Phan, and M. D. Wang, “Assessing the impact of human genome annotation choice on RNA-seq expression estimates,” Bmc Bioinformatics, vol. 14, Sep 13 2013.
[29] M. Tee, J. A. Noble, and D. A. Bluemke, “Imaging techniques for cardiac strain and deformation: comparison of echocardiography, cardiac magnetic resonance and cardiac computed tomography,” 2013.
[30] H. Wang and A. A. Amini, “Cardiac motion and deformation recovery from MRI: a review,” Medical Imaging, IEEE Transactions on, vol. 31, pp. 487-503, 2012.
[31] S. Cook, E. Ladich, G. Nakazawa, P. Eshtehardi, M. Neidhart, R. Vogel, M. Togni, P. Wenaweser, M. Billinger, and C. Seiler, “Correlation of intravascular ultrasound findings with histopathological analysis of thrombus aspirates in patients with very late drug-eluting stent thrombosis,” Circulation, vol. 120, pp. 391-399, 2009.
[32] K. P. Daly, A. C. Marshall, J. A. Vincent, W. A. Zuckerman, T. M. Hoffman, C. E. Canter, E. D. Blume, and L. Bergersen, “Endomyocardial biopsy and selective coronary angiography are low-risk procedures in pediatric heart transplant recipients: Results of a multicenter experience,” The Journal of Heart and Lung Transplantation, vol. 31, pp. 398-409, 2012.
[33] M. Mengel, B. Sis, D. Kim, J. Chang, K. Famulski, L. Hidalgo, G. Einecke, D. De Freitas, W. Tymchak, and J. Burton, “The molecular phenotype of heart transplant biopsies: relationship to histopathological and clinical variables,” American Journal of Transplantation, vol. 10, pp. 2105-2115, 2010.
[34] S. Kothari, J. H. Phan, T. H. Stokes, and M. D. Wang, “Pathology imaging informatics for quantitative analysis of whole-slide images,” Journal of the American Medical Informatics Association, vol. 20, pp. 1099-1108, 2013.
[35] S. Kothari, J. H. Phan, A. O. Osunkoya, and M. D. Wang, “Biological Interpretation of Morphological Patterns in Histopathological Whole-Slide Images,” in ACM Conference on Bioinformatics, Computational Biology and Biomedicine, 2012.
[36] S. Kothari, J. Phan, T. Stokes, A. Osunkoya, A. Young, and M. Wang, “Removing Batch Effects from Histopathological Images for Enhanced Cancer Diagnosis,” IEEE Journal of Biomedical and Health Informatics, vol. PP, p. 1.
[37] S. Kothari, J. H. Phan, A. N. Young, and M. D. Wang, “Histological image classification using biologically interpretable shape-based features,” BMC medical imaging, vol. 13, pp. 1-17, 2013.
[38] C. D. Kaddi, R. M. Parry, and M. D. Wang, “Multivariate Hypergeometric Similarity Measure,” IEEE-ACM Transactions on Computational Biology and Bioinformatics, vol. 10, pp. 1505-1516, Nov-Dec 2013.
[39] A. C. Grey, A. K. Gelasco, J. Section, R. A. Moreno-Rodriguez, E. L. Krug, and K. L. Schey, “Molecular morphology of the chick heart visualized by MALDI imaging mass spectrometry,” Anat Rec (Hoboken), vol. 293, pp. 821-8, May 2010.
[40] R. F. Menger, W. L. Stutts, D. S. Anbukumar, J. A. Bowden, D. A. Ford, and R. A. Yost, “MALDI mass spectrometric imaging of cardiac tissue following myocardial infarction in a rat coronary artery ligation model,” Anal Chem, vol. 84, pp. 1117-25, Jan 17 2012.
[41] R. M. Parry, A. S. Galhena, C. M. Gamage, R. V. Bennett, M. D. Wang, and F. M. Fernandez, “omniSpect: an open MATLAB-based tool for visualization and analysis of matrix-assisted laser desorption/ionization and desorption electrospray ionization mass spectrometry images,” J Am Soc Mass Spectrom, vol. 24, pp. 646-9, Apr 2013.
[42] L. A. Cooper, J. Kong, D. A. Gutman, F. Wang, S. R. Cholleti, T. C. Pan, P. M. Widener, A. Sharma, T. Mikkelsen, and A. E. Flanders, “An integrative approach for in silico glioma research,” Biomedical Engineering, IEEE Transactions on, vol. 57, pp. 2617-2621, 2010.
[43] H. Chang, G. V. Fontenay, J. Han, G. Cong, F. L. Baehner, J. W. Gray, P. T. Spellman, and B. Parvin, “Morphometic analysis of TCGA glioblastoma multiforme,” BMC bioinformatics, vol. 12, p. 484, 2011.
[44] C. Cheng, N. Chanani, J. Venugopalan, K. Maher, and M. D. Wang, “icuARM – An ICU Clinical Decision Support System Using Association Rule Mining,” IEEE J. Transl. Eng. Health Med., vol. 1, pp. 122-131, 2013.
[45] M. Saeed, M. Villarroel, A. T. Reisner, G. Clifford, L. W. Lehman, G. Moody, T. Heldt, T. H. Kyaw, B. Moody, and R. G. Mark, “Multiparameter Intelligent Monitoring in Intensive Care II: a public-access intensive care unit database,” Crit Care Med, vol. 39, pp. 952-60, May 2011.
Contributors
 Po-Yen Wu is a Ph.D. student in the School of Electrical and Computer Engineering at the Georgia Institute of Technology. His research interests include (1) designing novel methodologies to improve the data analysis pipeline for RNA-sequencing data, (2) exploring data-mining techniques for extracting meaningful patterns in electronic health records, and (3) integrating information from RNA-sequencing data and electronic health records to improve the prediction performance of clinical endpoints. Read more
Po-Yen Wu is a Ph.D. student in the School of Electrical and Computer Engineering at the Georgia Institute of Technology. His research interests include (1) designing novel methodologies to improve the data analysis pipeline for RNA-sequencing data, (2) exploring data-mining techniques for extracting meaningful patterns in electronic health records, and (3) integrating information from RNA-sequencing data and electronic health records to improve the prediction performance of clinical endpoints. Read more
 Janani Venugopalan is a Ph.D. student in the Joint Dept. of Biomedical Engineering at Georgia Institute of Technology and Emory University, Atlanta, Georgia. Her research focuses on health informatics. Her research focuses on clinical decision making using HER data, which are currently in collaboration with Children’s Healthcare of Atlanta and Emory. Read more
Janani Venugopalan is a Ph.D. student in the Joint Dept. of Biomedical Engineering at Georgia Institute of Technology and Emory University, Atlanta, Georgia. Her research focuses on health informatics. Her research focuses on clinical decision making using HER data, which are currently in collaboration with Children’s Healthcare of Atlanta and Emory. Read more
 Sonal Kothari received her Ph.D. degree in the School of Electrical and Computer Engineering at the Georgia Institute of Technology, Atlanta. She is currently a postdoctoral fellow working under Principle Investigator May D. Wang in the Department of Biomedical Engineering at Georgia Tech and Emory University. Her research interests are in the areas of imaging informatics, histopathological image analysis, pattern recognition, and visualization. Read more
Sonal Kothari received her Ph.D. degree in the School of Electrical and Computer Engineering at the Georgia Institute of Technology, Atlanta. She is currently a postdoctoral fellow working under Principle Investigator May D. Wang in the Department of Biomedical Engineering at Georgia Tech and Emory University. Her research interests are in the areas of imaging informatics, histopathological image analysis, pattern recognition, and visualization. Read more
 Chanchala D. Kaddi is currently a Ph.D. student in Bioengineering at Georgia Institute of Technology. She is a National Science Foundation Graduate Research Fellow. Her research interests are in computational systems biology and bioinformatics, including methods for mass spectrometry imaging data analysis and predictive models for cancer research. Read more
Chanchala D. Kaddi is currently a Ph.D. student in Bioengineering at Georgia Institute of Technology. She is a National Science Foundation Graduate Research Fellow. Her research interests are in computational systems biology and bioinformatics, including methods for mass spectrometry imaging data analysis and predictive models for cancer research. Read more
 Chih-Wen Cheng is currently pursuing his Ph.D. as a graduate research assistant in the School of Electrical and Computer Engineering. His research focuses on applying data mining, cloud computing, and mobile technologies to improve clinical decision-making to facilitate quality-of-healthcare. Read more
Chih-Wen Cheng is currently pursuing his Ph.D. as a graduate research assistant in the School of Electrical and Computer Engineering. His research focuses on applying data mining, cloud computing, and mobile technologies to improve clinical decision-making to facilitate quality-of-healthcare. Read more
 John H. Phan is an assistant research professor in the Department of Biomedical Engineering, Georgia Institute of Technology and Emory University, Atlanta, Georgia, USA. He received his Ph.D. degree in biomedical engineering from the Georgia Institute of Technology and Emory University. His research is focused on knowledge-driven high-throughput analysis of biomedical data, high-performance and grid computing, and integrative bioinformatics. Read more
John H. Phan is an assistant research professor in the Department of Biomedical Engineering, Georgia Institute of Technology and Emory University, Atlanta, Georgia, USA. He received his Ph.D. degree in biomedical engineering from the Georgia Institute of Technology and Emory University. His research is focused on knowledge-driven high-throughput analysis of biomedical data, high-performance and grid computing, and integrative bioinformatics. Read more
 May Wang is an Associate Professor in the Joint Department of BME, School of ECE, The Winship Cancer Institute, IBB, and IPaT at Georgia Institute of Technology and Emory University, USA. She is a Georgia Research Alliance Distinguished Cancer Scholar, Director of the Biocomputing and Bioinformatics Core in Emory-GeorgiaTech Cancer Nanotechnology Center, and Co-Director of Georgia-Tech Center of Bio-Imaging Mass Spectrometry. Prof. Wang’s research is on Big Biomedical Data Analytics with a focus on Biomedical and Health Informatics (BHI) for Personalized and Predictive Health. It includes high throughput next generation sequencing and -omic data mining to identify clinical biomarkers, pathological imaging informatics, health informatics, bionanoinformatics, and systems modeling. Read more
May Wang is an Associate Professor in the Joint Department of BME, School of ECE, The Winship Cancer Institute, IBB, and IPaT at Georgia Institute of Technology and Emory University, USA. She is a Georgia Research Alliance Distinguished Cancer Scholar, Director of the Biocomputing and Bioinformatics Core in Emory-GeorgiaTech Cancer Nanotechnology Center, and Co-Director of Georgia-Tech Center of Bio-Imaging Mass Spectrometry. Prof. Wang’s research is on Big Biomedical Data Analytics with a focus on Biomedical and Health Informatics (BHI) for Personalized and Predictive Health. It includes high throughput next generation sequencing and -omic data mining to identify clinical biomarkers, pathological imaging informatics, health informatics, bionanoinformatics, and systems modeling. Read more






 Mary Capelli-Schellpfeffer, MD, MPA, is Medical Director of Loyola University Health System's Occupational Health Services, and Associate Professor, Department of Medicine, Loyola University Chicago Stritch School of Medicine. Dr. Mary Capelli-Schellpfeffer guides Loyola's occupational medicine programs. An IEEE Fellow and global consultant regarding electrical safety, she is widely recognized for her work on risk mitigation in complex systems.
Mary Capelli-Schellpfeffer, MD, MPA, is Medical Director of Loyola University Health System's Occupational Health Services, and Associate Professor, Department of Medicine, Loyola University Chicago Stritch School of Medicine. Dr. Mary Capelli-Schellpfeffer guides Loyola's occupational medicine programs. An IEEE Fellow and global consultant regarding electrical safety, she is widely recognized for her work on risk mitigation in complex systems.  Emily Sopensky co-founded the RFID in Healthcare Consortium in 2008 as an educational not-for-profit addressing technology primarily in healthcare facilities. Initially created 5 years ago by the RHCC, 2014 is the first year the Symposium has substantial IEEE involvement (Life Sciences and Engineering Medicine and Biology Society). As an active IEEE volunteer and Chair-Elect, IEEE Technical Committee on RFID (CRFID), she anticipates next year's symposium will similarly have active IEEE participation.
Emily Sopensky co-founded the RFID in Healthcare Consortium in 2008 as an educational not-for-profit addressing technology primarily in healthcare facilities. Initially created 5 years ago by the RHCC, 2014 is the first year the Symposium has substantial IEEE involvement (Life Sciences and Engineering Medicine and Biology Society). As an active IEEE volunteer and Chair-Elect, IEEE Technical Committee on RFID (CRFID), she anticipates next year's symposium will similarly have active IEEE participation.  Kuldeep Singh V Rajput is a Project Associate at the Indian Institute of Technology, Madras, India. He received the B. Tech degree from the B.V. Bhoomaraddi College of Engineering and Technology. His research interests are in the field of VLSI, Computer Vision and Graphics, Haptics, Signal Processing, Biomedical Engineering and Biosensors.
Kuldeep Singh V Rajput is a Project Associate at the Indian Institute of Technology, Madras, India. He received the B. Tech degree from the B.V. Bhoomaraddi College of Engineering and Technology. His research interests are in the field of VLSI, Computer Vision and Graphics, Haptics, Signal Processing, Biomedical Engineering and Biosensors.  Po-Yen Wu is a Ph.D. student in the School of Electrical and Computer Engineering at the Georgia Institute of Technology. His research interests include (1) designing novel methodologies to improve the data analysis pipeline for RNA-sequencing data, (2) exploring data-mining techniques for extracting meaningful patterns in electronic health records, and (3) integrating information from RNA-sequencing data and electronic health records to improve the prediction performance of clinical endpoints.
Po-Yen Wu is a Ph.D. student in the School of Electrical and Computer Engineering at the Georgia Institute of Technology. His research interests include (1) designing novel methodologies to improve the data analysis pipeline for RNA-sequencing data, (2) exploring data-mining techniques for extracting meaningful patterns in electronic health records, and (3) integrating information from RNA-sequencing data and electronic health records to improve the prediction performance of clinical endpoints.  Janani Venugopalan is a Ph.D. student in the Joint Dept. of Biomedical Engineering at Georgia Institute of Technology and Emory University, Atlanta, Georgia. Her research focuses on health informatics. Her research focuses on clinical decision making using HER data, which are currently in collaboration with Children's Healthcare of Atlanta and Emory.
Janani Venugopalan is a Ph.D. student in the Joint Dept. of Biomedical Engineering at Georgia Institute of Technology and Emory University, Atlanta, Georgia. Her research focuses on health informatics. Her research focuses on clinical decision making using HER data, which are currently in collaboration with Children's Healthcare of Atlanta and Emory.  Sonal Kothari received her Ph.D. degree in the School of Electrical and Computer Engineering at the Georgia Institute of Technology, Atlanta. She is currently a postdoctoral fellow working under Principle Investigator May D. Wang in the Department of Biomedical Engineering at Georgia Tech and Emory University. Her research interests are in the areas of imaging informatics, histopathological image analysis, pattern recognition, and visualization.
Sonal Kothari received her Ph.D. degree in the School of Electrical and Computer Engineering at the Georgia Institute of Technology, Atlanta. She is currently a postdoctoral fellow working under Principle Investigator May D. Wang in the Department of Biomedical Engineering at Georgia Tech and Emory University. Her research interests are in the areas of imaging informatics, histopathological image analysis, pattern recognition, and visualization.  Chanchala D. Kaddi is currently a Ph.D. student in Bioengineering at Georgia Institute of Technology. She is a National Science Foundation Graduate Research Fellow. Her research interests are in computational systems biology and bioinformatics, including methods for mass spectrometry imaging data analysis and predictive models for cancer research.
Chanchala D. Kaddi is currently a Ph.D. student in Bioengineering at Georgia Institute of Technology. She is a National Science Foundation Graduate Research Fellow. Her research interests are in computational systems biology and bioinformatics, including methods for mass spectrometry imaging data analysis and predictive models for cancer research.  Chih-Wen Cheng is currently pursuing his Ph.D. as a graduate research assistant in the School of Electrical and Computer Engineering. His research focuses on applying data mining, cloud computing, and mobile technologies to improve clinical decision-making to facilitate quality-of-healthcare.
Chih-Wen Cheng is currently pursuing his Ph.D. as a graduate research assistant in the School of Electrical and Computer Engineering. His research focuses on applying data mining, cloud computing, and mobile technologies to improve clinical decision-making to facilitate quality-of-healthcare.  John H. Phan is an assistant research professor in the Department of Biomedical Engineering, Georgia Institute of Technology and Emory University, Atlanta, Georgia, USA. He received his Ph.D. degree in biomedical engineering from the Georgia Institute of Technology and Emory University. His research is focused on knowledge-driven high-throughput analysis of biomedical data, high-performance and grid computing, and integrative bioinformatics.
John H. Phan is an assistant research professor in the Department of Biomedical Engineering, Georgia Institute of Technology and Emory University, Atlanta, Georgia, USA. He received his Ph.D. degree in biomedical engineering from the Georgia Institute of Technology and Emory University. His research is focused on knowledge-driven high-throughput analysis of biomedical data, high-performance and grid computing, and integrative bioinformatics.  May Wang is an Associate Professor in the Joint Department of BME, School of ECE, The Winship Cancer Institute, IBB, and IPaT at Georgia Institute of Technology and Emory University, USA. She is a Georgia Research Alliance Distinguished Cancer Scholar, Director of the Biocomputing and Bioinformatics Core in Emory-GeorgiaTech Cancer Nanotechnology Center, and Co-Director of Georgia-Tech Center of Bio-Imaging Mass Spectrometry. Prof. Wang's research is on Big Biomedical Data Analytics with a focus on Biomedical and Health Informatics (BHI) for Personalized and Predictive Health. It includes high throughput next generation sequencing and -omic data mining to identify clinical biomarkers, pathological imaging informatics, health informatics, bionanoinformatics, and systems modeling.
May Wang is an Associate Professor in the Joint Department of BME, School of ECE, The Winship Cancer Institute, IBB, and IPaT at Georgia Institute of Technology and Emory University, USA. She is a Georgia Research Alliance Distinguished Cancer Scholar, Director of the Biocomputing and Bioinformatics Core in Emory-GeorgiaTech Cancer Nanotechnology Center, and Co-Director of Georgia-Tech Center of Bio-Imaging Mass Spectrometry. Prof. Wang's research is on Big Biomedical Data Analytics with a focus on Biomedical and Health Informatics (BHI) for Personalized and Predictive Health. It includes high throughput next generation sequencing and -omic data mining to identify clinical biomarkers, pathological imaging informatics, health informatics, bionanoinformatics, and systems modeling.