Single-cell Genome Sequencing and Assembly: Progress and Prospects
By H. Chitsaz
Recent technological advances in high throughput low-cost DNA sequencing have brought a whole new realm of exciting applications within reach, one of which is genomic analysis at single-cell resolution. Single-cell genome sequencing holds great promise for environmental biology. New techniques for single-cell sequencing are described.
Recent technological advances in high throughput low-cost DNA sequencing have brought a whole new realm of exciting applications within reach, one of which is genomic analysis at single-cell resolution. Single-cell genome sequencing holds great promise for environmental biology. The vast majority of environmental bacteria, e.g. from human microbiome, cannot be cultivated in isolation as they require their symbiotic natural habitat to grow. Also, single-cell genome sequencing holds great promise for various other areas of biology, e.g. cancer research. It is well known that a cancerous tumor is heterogeneous as the constituent cells quickly accumulate random mutations and aberrations in the absence of gatekeeper (watchdog) genes. An important problem in cancer genomics is discovery of the first few initial variations that started the tumor. A solution to that problem consists in reconstruction of the tumor phylogeny from the genomes of single cells sampled from different parts of the tumor. Therefore, our ability to acquire and analyze genomic sequences at single-cell resolution is expected to have significant impact on energy, environmental, and health research.
How does genome sequencing and assembly work? Whole genome shotgun sequencing starts with multiple copies of the genomic DNA. Each DNA molecule is broken into short fragments (100-10,000 bps), at random, which are then fed into a DNA sequencing machine to read their nucleotide sequence. The output reads are short randomly sampled subsequences of the genomic sequence. Since multiple copies of the genomic DNA are sequenced, every genomic locus is covered by multiple reads, whose number is called coverage. If the coverage is sufficient, then reads can be concatenated along their suffix-prefix overlaps to form longer contiguous genomic subsequences (contigs) in a de novo assembly algorithm [1]. The number of reads in a human genome sequencing experiment with Illumina HiSeq platform, for instance, can easily reach 1.5 billion. Hence, de novo assembly is essentially like solving a gigantic jigsaw puzzle which is stained by sequencing errors and obscured by repetitive elements, e.g. genomic Alu repeats and segmental duplications.
A single cell contains only one DNA molecule whereas whole genome shotgun sequencing requires multiple copies. Fortunately, this gap can now be filled by DNA amplification techniques that replicate an input DNA template up to several billion folds. A popular choice for whole genome amplification is currently multiple displacement amplification (MDA) [2][9]. However, DNA amplification comes with inconvenient side effects including chimeric junctions in the amplicons and largely biased amplification gains (Figure 1). In this case, differentiating a low-coverage correct contig from an erroneous or chimeric piece of sequence becomes a challenge. We developed a special de novo assembly algorithm to deal with such problems caused by DNA amplification [3]. More advanced algorithms, e.g. SPAdes [7] and IDBA-UD [8], were introduced later to rescue more low-coverage correct contigs.
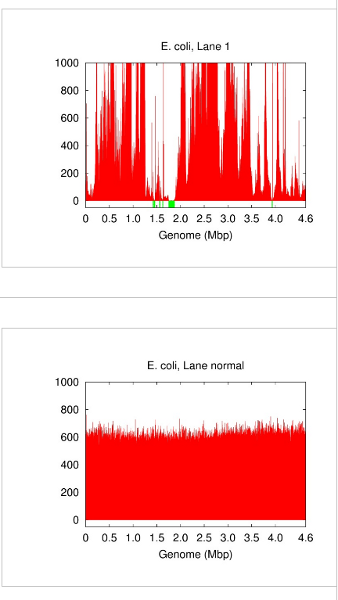
Figure 1: Genome coverage in single-cell E. coli (lane 1 in [3]) vs. normal multicell E. coli. Both have an average coverage of ~600x. Blackout (no-coverage) regions are shown in green.
Although we demonstrated that single-cell sequencing methods have passed important milestones, e.g. capturing > 90% of genes in a prokaryotic cell [3], their quality and reliability linger unsurprisingly behind those of normal multicell sequencing methods. Beside sporadic gaps because of lack of coverage and chimeric contigs, the main practical problem with single-cell sequencing is wide variability of the outcome. On one side of the spectrum, the genome can be almost completely acquired. On the other side, however, the outcome may very well be the loss of the sample and any information therein. In this sense, a single-cell sequencing experiment is currently a gamble that can potentially lead to the loss of the sample and sequencing expenses. Therefore, a few single cells (e.g. 10-20 cells) are isolated and sequenced in practice to hedge against that risk.
A mechanism to detect and enrich target cells is required in that approach. Isolation of a number of cells from the same species would be costly and even impossible when there is no prior knowledge about the organism. A solution that we proposed replaces a single-cell deep sequencing experiment with numerous single-cell shallow sequencing experiments using less stringent isolation or enrichment, leaving the total cost of sequencing intact [4]. That approach hedges against the risk of total loss of information based on the observation that DNA amplification bias is not sequence-specific, meaning that the same genomic loci may be amplified poorly in one amplification reaction and adequately in another. We used this characteristic to decrease the coverage bias and increase assembly quality through mixing reads from few individual single cells. To avoid mixing reads from non-identical genomes, which would create a chimeric assembly, our de novo co-assembly algorithm HyDA assigns a unique color to each input single-cell read data set and keeps track of the color of each read and contig in the assembly process. In the end, contigs of every color are separately presented in the output. Our algorithm identifies those contigs that are exclusive to a color and those that are shared between two or more colors. That information is used to construct a belief state about the relationships between colors, which in turn is used to cluster those colors that are believed to be from the same species [4].
That method requires a less stringent but nevertheless a mechanism for enrichment of target cells. What if there is absolutely no prior knowledge about the target organism(s)? In that case, sequencing, assembly, and identification of every distinct genome (species) in the sample would be desired. That is challenging as there are typically millions, sometimes billions, of cells in a microbial sample [5]. Exhaustive single-cell sequencing is simply intractable. However, there is hope for breaking that barrier as the number of different species is usually much smaller than the number of cells. We call this feature species sparsity in a sample. We gave a novel divide-and-conwuer algorithm to sequence and de novo assemble all distinct genomes in a microbial sample with reduced sequencing cost and computational effort [6]. Our method is implemented in a tool called Squeezambler, which is packaged with HyDA as an opensource toolbox available at http://sourceforge.net/projects/hyda/.
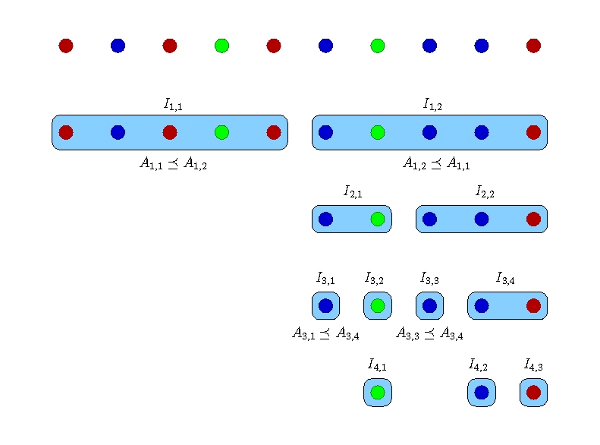
Figure 2: The divide-and-conwuer algorithm for an example with 10 cells and 3 distinct genomes shown in different colors [6]. Each row corresponds to one sequencing round. The number of barcodes in each round is the number of blue boxes in the corresponding row.
In the divide-and-conwuer method, the DNA of every cell is individually amplified and sequenced at adaptive depths to reduce redundant sequencing of one species in numerous cells. In this process (Figure 2), a search tree (explained below) is constructed, and the technique to make and traverse the tree has an important role in determining the amount of resources to be allocated at each step of the algorithm and the overall sequencing cost. Let n be the number of cells in the sample, and denote the cells by Si, i = 1,…,n. In the first iteration, the algorithm divides the n cells into two sets I1,1= {S1,…,S[n/2]}, I1,2= {S[n/2]+1,…,Sn}. Our algorithm samples equal amount of amplicons from each cell in I1,1 and I1,2. The amplicons in each set are pooled and sequenced to reach a desired number of base pairs that is computed by the algorithm depending on the number of cells in the pool and an estimate of the total assembly size when possible. In practice, the two pools are multiplexed by two distinct tag barcodes and sequenced in one run of a high throughput sequencing machine. The barcoded amplicons are sequenced, and the two read datasets are coassembled by HyDA using two colors. The result is two sets of contigs, one for each color, A1,1 and A1,2. Based on the exclusivity of A1,1 and A1,2, we decide if one set subsumes the other, i.e. A1,1 ≤ A1,2 or A1,2 ≤ A1,1. For instance, if A1,1 is subsumed by A1,2, then all of the distinct genomes in I1,1 are present in I1,2; therefore, the cells in I1,1 are redundant and do not need further sampling. If neither set subsumes the other, both I1,1 and I1,2 remain for the next step. Each remaining set I1,* is divided into two subsets for analysis in the second iteration. The same splitting process occurs in the subsequent iterations until each remaining set contains only one cell. Figure 2 depicts an example of 10 cells with 3 distinct genomes shown in different colors [6].
In spite of significant progress, the field of single-cell genome sequencing and assembly is still open and expanding. Particularly, whole sample approaches, the first of which was our adaptive divide-and-conwuer algorithm [6], are expected to be rich enough to breed a subfield of single-cell sequencing.
Acknowledgement
The research works mentioned here were in collaboration with Jonathan H. Badger, Sorin Drăghici, Christopher L. Dupont, Dirk J. Evers, Louise J. Fraser, Niall A. Gormley, Roger S. Lasken, Mary-Jane Lombardo, Narjes S. Movahedi Tabrizi, Mark Novotny, Pavel A. Pevzner, Douglas B. Rusch, Ole Schulz-Trieglaff, Geoffrey P. Smith, Zeinab Taghavi, Glenn Tesler, Joyclyn L. Yee-Greenbaum, and supported partially by NSF DBI-1262565.
For Further Reading
1. Compeau, P.E.C. et al. “How to apply de Bruijn graphs to genome assembly”. Nature Biotechnology, 29 (10), 987-991 (2011).
2. Dean, F.B. et al. “Comprehensive human genome amplification using multiple displacement amplification”. Proc. Natl. Acad. Sci. USA 99, 5261-5266 (2002).
3. Hamidreza Chitsaz, Joyclyn L. Yee-Greenbaum, Glenn Tesler, MaryJane Lombardo, Christopher L. Dupont, Jonathan H. Badger, Mark Novotny, Douglas B. Rusch, Louise J. Fraser, Niall A. Gormley, Ole Schulz-Trieglaff, Geoffrey P. Smith, Dirk J. Evers, Pavel A. Pevzner, and Roger S. Lasken. “Efficient de novo assembly of single-cell bacterial genomes from short-read data sets”. Nature Biotechnology, 29 (10), 915-921, (2011).
4. Narjes S. Movahedi, Elmirasadat Forouzmand, and Hamidreza Chitsaz. “De Novo Co-assembly of Bacterial Genomes from Multiple Single Cells”. IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 561-565 (2012).
5. Qin J, et al. “A human gut microbial gene catalogue established by metagenomic sequencing”. Nature 464, 59-65 (2010).
6. Zeinab Taghavi, Narjes S. Movahedi, Sorin Drăghici, and Hamidreza Chitsaz. “Distilled single-cell genome sequencing and de novo assembly for sparse microbial communities”. Bioinformatics 29 (19), 2395-2401 (2013).
7. Bankevich A, et al. “SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing”. J. Comput. Biol. 19, 455-477 (2012).
8. Peng Y, et al. “IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth”. Bioinformatics 28, 1420-1428 (2012).
9. Dean, F.B., Nelson, J.R., Giesler, T.L. & Lasken, R.S. “Rapid amplification of plasmid and phage DNA using Phi 29 DNA polymerase and multiply-primed rolling circle amplification”. Genome Res. 11, 1095-1099 (2001).
Contributor
 H. Chitsaz is assistant professor of Computer Science at Wayne State University, Detroit, MI. He received his Ph.D. in Computer Science and M.S. in Mathematics from the University of Illinois at Urbana-Champaign. Read more
H. Chitsaz is assistant professor of Computer Science at Wayne State University, Detroit, MI. He received his Ph.D. in Computer Science and M.S. in Mathematics from the University of Illinois at Urbana-Champaign. Read more






 Dr. Stephen Quake, D.Phil., is a Professor of Bioengineering and of Applied Physics at Stanford University. He pioneered the development of Microfluidic Large Scale Integration (LSI), demonstrating the first integrated microfluidic devices with thousands of mechanical valves.
Dr. Stephen Quake, D.Phil., is a Professor of Bioengineering and of Applied Physics at Stanford University. He pioneered the development of Microfluidic Large Scale Integration (LSI), demonstrating the first integrated microfluidic devices with thousands of mechanical valves.  H. Chitsaz is assistant professor of Computer Science at Wayne State University, Detroit, MI. He received his Ph.D. in Computer Science and M.S. in Mathematics from the University of Illinois at Urbana-Champaign.
H. Chitsaz is assistant professor of Computer Science at Wayne State University, Detroit, MI. He received his Ph.D. in Computer Science and M.S. in Mathematics from the University of Illinois at Urbana-Champaign.  Mathukumalli Vidyasagar is the Founding Head of the Bioengineering Department, University of Texas at Dallas. He is a Fellow of the Royal Society, UK.
Mathukumalli Vidyasagar is the Founding Head of the Bioengineering Department, University of Texas at Dallas. He is a Fellow of the Royal Society, UK.  Usmah Kawoos (S '05-M '09) received the Ph.D degree in biomedical engineering from Drexel University, Philadelphia, PA. Dr. Kawoos is currently working as a post-doctoral fellow at the Naval Medical Research Center, Silver Spring, MD.
Usmah Kawoos (S '05-M '09) received the Ph.D degree in biomedical engineering from Drexel University, Philadelphia, PA. Dr. Kawoos is currently working as a post-doctoral fellow at the Naval Medical Research Center, Silver Spring, MD.  Xu Meng (S '08-M '13) received the Ph.D. degree in biomedical engineering in 2013 from Drexel University, Philadelphia, PA. His research interests are the clinical application of microwave and RF in medical implants and telemedicine.
Xu Meng (S '08-M '13) received the Ph.D. degree in biomedical engineering in 2013 from Drexel University, Philadelphia, PA. His research interests are the clinical application of microwave and RF in medical implants and telemedicine.  Arye Rosen (M '77-SM '80-F '92) received the Ph.D. degree in electrical engineering from Drexel University, Philadelphia, PA. Dr. Rosen currently holds an appointment as Academy Professor of Biomedical and Electrical Engineering in the School of Biomedical Engineering, Science and Health Systems and Associate Vice Provost for Strategic Initiatives at Drexel University, Philadelphia, PA.
Arye Rosen (M '77-SM '80-F '92) received the Ph.D. degree in electrical engineering from Drexel University, Philadelphia, PA. Dr. Rosen currently holds an appointment as Academy Professor of Biomedical and Electrical Engineering in the School of Biomedical Engineering, Science and Health Systems and Associate Vice Provost for Strategic Initiatives at Drexel University, Philadelphia, PA.  Mikulas Chavko received the PhD degree in Biochemistry from University of Veterinary Medicine in Kosice, Czechoslovakia. Currently, he serves as a Principal Investigator and Head of Blast Research in the Neurotrauma Department at the Naval Medical Research Institute.
Mikulas Chavko received the PhD degree in Biochemistry from University of Veterinary Medicine in Kosice, Czechoslovakia. Currently, he serves as a Principal Investigator and Head of Blast Research in the Neurotrauma Department at the Naval Medical Research Institute.