Computational Biology Corner
By Mathukumalli Vidyasagar
In his continuing column, this month Dr. Vidyasagar discusses how the variability of biological data, derived from different commercial platforms, impedes the application of engineering methodology.
As I spend more and more time grappling with biological data sets, trying to apply methods of engineering (and mathematical) modeling and analysis, I am struck by how much variability there is in the data sets. By this I do not mean the enormous diversity of biology as a subject, and the resulting diversity of the data. Rather, I refer to the fact that many biological experiments are not fully repeatable, and thus the resulting data sets are not readily amenable to the application of methods that people like us take for granted.
A rather dramatic report on the non-repeatability of biological experiments is to be found in a recent editorial [1] in Nature by C. Glenn Begley, formerly Global Head of Hematology and Oncology Research at Amgen, and Lee M. Ellis of the University of Texas M. D. Anderson Cancer Center. It is claimed in the editorial that when Dr. Begley asked his team at Amgen to reproduce the findings of 53 “landmark” papers in this area, only six results could be reproduced, and by implication 47 results could not be reproduced. As the authors did not state which 53 papers were so tested, this has led to predictable jokes about the findings of the editorial themselves not being reproducible. Leaving aside that issue however, I can readily sympathize with these authors, as I have had similar experiences.
Rather than comment in vague generalities and broad-brush strokes, let me focus on an issue that has vexed me during recent months, namely: platform dependence of data. As the name suggests, “platform dependence” refers to the phenomenon whereby multiple platforms from commercial vendors, all of them purporting to measure exactly the same quantity, produce results that bear no resemblance to each other. The biology community uses the term “platform” as we use “apparatus”. The specific platforms with which I am currently struggling are those for measuring gene expression data. An excellent review of the technology involved is found in [2].
Let me be even more specific. TCGA (The Cancer Genome Atlas) is a very ambitious project undertaken by the National Cancer Institute, to derive molecular information about (eventually) every cancer tissue that is available. In a recent paper [2], the TCGA Research Network reported its findings on ovarian cancer. Nearly 600 cancer tissues surgically removed from patients were analyzed using two popular commercial platforms, both of them beginning with the letter A. Expression levels of nearly 20,000 genes in one case and nearly 12,000 genes in the other case were measured using the respective platforms. One of the challenges in ovarian cancer is that about 20% of the patients simply do not respond to standard front-line therapy, which consists of administering platinum with taxane, or cisplatin, or some such drug. It would therefore be highly desirable if one could determine, on the basis of the expression levels of a small number of genes from the tumor, whether the patient is unlikely to respond. If such a prediction could be made with high accuracy, the treating physician could have a back-up plan ready for those who are not likely to respond to platinum chemotherapy. My students and I had developed a promising algorithm for doing precisely this, and tested it on expression levels measured from Platform A. The results were simply spectacular, with accuracies in excess of 90%. Buoyed by success, and heedless of the pitfalls ahead, we then rather rashly applied our classifier to data from Platform B (or perhaps I should call it Platform A’, as its name also begins with A). To say that the performance of the classifier was mediocre is to put it mildly. We checked our equations, and then our code, and only then did we think of comparing the measurements from the two platforms.
The three figures below show the measurements of the expression level of the same gene on the two platforms, across multiple tumor tissues. Figure 1 for the gene CARTPT, Figure 2 is for TRIM15 and Figure 3 is for C11orf76. The plot for CARTPT looks more or less linear, so that is acceptable. The plot for TRIM15 is not so coherent, but perhaps an exponential plot might fit the data reasonably well. But what are we to make of the plot for C11orf76, which resembles pure noise as much as anything?
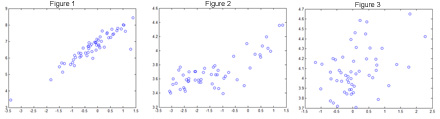
Figure 1: Measurements of the Expression Levels of CARTPT.
Figure 2: Measurements of the Expression Levels of TRIM15.
Figure 3: Measurements of the Expression Levels of C110rf76.
The difficulties arise because, while both platforms are supposed to be measuring the same thing, in reality they use quite different “probes”. In engineering we are quite accustomed to one instrument being replaced or coexisting with another one to measure the same quantity. But I am hard-pressed to find an instance in engineering where two instruments produce such wildly divergent measurements of the same quantity.
It is not that biologists are unaware of these phenomena. Given the horrendous complexity of biological systems, it is a near-miracle that one is able to measure anything at such a microscopic scale. When I discussed the contents of this column with my collaborator, Prof. Michael A. White of the famed UT Southwestern Medical Center in Dallas, he commented that “Technology industry [in biology] off-loads quality control to the users. We do a lot of pro bono quality control for industry”. My reaction was that in engineering we don’t do pro bono quality control or fault identification (unless the products in question are from Microsoft). Therefore, when we keep hearing about biological data being generated at an unprecedented pace, and how now is the right time for us engineers to jump into computational biology, we would do well to ask ourselves what it is that we can really offer when the raw data sets are so non-repeatable. At the very least, we should avoid the hubris of thinking that methods that are well-established in engineering settings would work equally well in biological problems.
For Further Reading
-
C. Glenn Begley and Lee M. Ellis, “Drug development: Raise standards for preclinical cancer research”, Nature, 483, 531-533, 29 March 2012.
-
M. Madan Babu, “An introduction to microarray data analysis”, in Computational Genomics: Theory and Applications, Horizon Bioscience, 2004.
-
The Cancer Genome Atlas Research Network, Nature, 474, 609-615, 30 June 2011.






 Thomas Schmitz-Rode is Chairman, Applied Medical Engineering at RWTH Aachen University. He received a Dipl.-Ing. degree in Mechanical Engineering from RWTH Aachen University in 1982.
Thomas Schmitz-Rode is Chairman, Applied Medical Engineering at RWTH Aachen University. He received a Dipl.-Ing. degree in Mechanical Engineering from RWTH Aachen University in 1982.  Atam P. Dhawan, Ph.D., obtained his bachelor's and master's degrees from the Indian Institute of Technology, Roorkee, and Ph.D. from the University of Manitoba, in Electrical Engineering.
Atam P. Dhawan, Ph.D., obtained his bachelor's and master's degrees from the Indian Institute of Technology, Roorkee, and Ph.D. from the University of Manitoba, in Electrical Engineering.  Mathukumalli Vidyasagar received B.S., M.S. and Ph.D. degrees in electrical engineering from the University of Wisconsin in Madison, in 1965, 1967 and 1969 respectively.
Mathukumalli Vidyasagar received B.S., M.S. and Ph.D. degrees in electrical engineering from the University of Wisconsin in Madison, in 1965, 1967 and 1969 respectively.